Javascript API¶
Overview¶
Embedding HiGlass in web page¶
HiGlass can be included in any web page by including the relevant javascript and css files:
<!DOCTYPE html>
<head>
<meta charset="utf-8">
<link rel="stylesheet" href="https://unpkg.com/higlass@1.5.7/dist/hglib.css" type="text/css">
<link rel="stylesheet" href="//maxcdn.bootstrapcdn.com/bootstrap/3.3.7/css/bootstrap.min.css" type="text/css">
<script crossorigin src="https://unpkg.com/react@16.6/umd/react.production.min.js"></script>
<script crossorigin src="https://unpkg.com/react-dom@16.6/umd/react-dom.production.min.js"></script>
<script crossorigin src="https://unpkg.com/pixi.js@5/dist/pixi.min.js"></script>
<!-- To render HiGlass with the Canvas API include the pixi.js-legacy instead of pixi.js -->
<!-- <script crossorigin src="https://unpkg.com/pixi.js-legacy@5/dist/pixi-legacy.min.js"></script> -->
<script src="https://cdnjs.cloudflare.com/ajax/libs/react-bootstrap/0.32.1/react-bootstrap.min.js"></script>
<script src="https://unpkg.com/higlass@1.6/dist/hglib.min.js"></script>
</head>
<body >
<div id="development-demo" style="width: 800px;
background-color: white;"></div>
</body>
<script>
const hgApi = hglib.viewer(
document.getElementById('development-demo'),
'http://higlass.io/api/v1/viewconfs/?d=default',
{
bounded: false,
}
);
</script>
</html>
External tracks should be included before the hglib.js import:
<script src="https://unpkg.com/higlass-multivec@0.2.0/dist/higlass-multivec.js"></script>
Instructions for instantiating the component and interacting with it are in the Public API section.
Available endpoints¶
import { HiGlassComponent, ChromosomeInfo, viewer, version } from 'higlass';
import 'higlass/dist/hglib.css';
HiGlass exports four endpoints for your convenience. viewer is the main
endpoint to create a new HiGlass component. HiGlassComponent can be used
to integrate HiGlass in your React application. ChromosomeInfo is a class
for converting absolute coordinates to chromosome coordinates. It’s used
internally and made available to convert absolute range selection into
chromosome range selections. version is a string of the current version of
HiGlass.
In order to get the expected look and feel, we also import the HiGlass CSS files. (Make sure your bundler knows how to handle CSS imports! E.g., if you’re using webpack you may need to include the [css-loader]( https://webpack.js.org/loaders/css-loader/).)
Creating an inline HiGlass component¶
const hgv = hglib.viewer(element, config, options);
Create a new HiGlass viewer within a web page. This initializes a
HiGlassComponent inside the element element with a viewconfig passed in as
config. If config is a string, it is interpreted as a url and used to
try to fetch a remote viewconfig.
The options parameter can have the following properties:
bounded: tells the HiGlass component to fill all the space in the containing element. Note that ifboundedis set to true, thenelementmust have a fixed heightpixelPreciseMarginPadding: iftrueapply pixel precise view height, padding, and margin.containerPaddingXandcontainerPaddingY: x and y padding react grid layout containers. The x padding resembles left and right padding of the entire react grid layout container, i.e., it can be interpreted as the global padding of an HiGlass instance. The y padding stands for the top and bottom padding but in case that the HiGlass view is not bound it will only add padding to the top. You can find out more about the container padding at https://github.com/STRML/react-grid-layout#grid-layout-props.viewMarginTop,viewMarginBottom,viewMarginLeft, andviewMarginRight: top, bottom, left, right margin between views in pixels. The margin area is not interactive, i.e., dragging on the margin area will not change the location of the view!viewPaddingTop,viewPaddingBottom,viewPaddingLeft, andviewPaddingRight: top, bottom, left, right padding between views in pixels. The padding area is interactive, i.e., dragging on the margin area will change the location of the view!broadcastMousePositionGlobally: iftruethe relative mouse position of this HiGlass instances (in data coordinates) will be broadcasted globally. This allows you to show the global mouse position in another HiGlass instance within the same browser tab or another browser tab.showGlobalMousePosition: iftrueany globally broadcasted mouse position will be shown for all tracks that haveoptions.showMousePosition = true.globalMousePosition: iftruethis will turn onbroadcastMousePositionGloballyandshowGlobalMousePosition. This is basically a convenience option to quickly broadcast and show global mouse positions.PIXI: Use a different PIXI library. Useful if trying to use a canvas renderer. Example:
import * as PIXI from "pixi.js-legacy"
<HiGlassComponent
options={
renderer: 'canvas',
PIXI
}
/>
renderer: ifcanvasHiGlass will render to the Canvas API. Otherwise
it will use WebGL. Need to pass in a legacy PIXI import as well. See the PIXI parameter above.
sizeMode: the size mode determines the visible height of the HiGlass instance. There are 4 modes: 1.default: the height is given by the sum of the tracks’ heights 2.bounded: tells the HiGlass component to bind the height to the parent container by dynamically adjusting the height of center tracks. 3.scroll: will activate scrolling by stretching HiGlass’ drawing surface to the extent ofelementand hiding overflowing content in the x direction and allowing to scroll when content overflows in the y direction. This mode will also set all views to zoom fixed automatically so that you are not scrolling and zooming at the same time. 4.overflow: same asscrollexcept that you can’t scroll. This mode is only needed when you want to dynamically switch between scrolling and pan+zooming. Say you scrolled halfway down and then want to temporarily pan&zoom a track at that position. If you would switch back toboundedthe scrollTop position would be lost becauseboundeddemands that your entire view is bound to the parent. Instead you want can switch tooverflowto keep the current scrollTop position and enable pan&zooming.Visually you can think of the four size modes as follows:
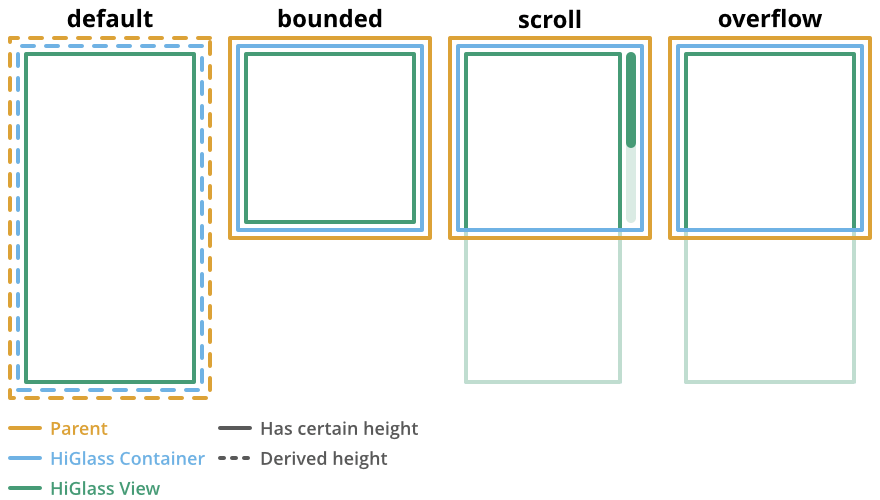
Note that if
sizeModeis anything other thandefault, theelementmust have a fixed height!
The function returns an instance of the public API of a HiGlass component.
A full example of an inline HiGlass component can be found in the HiGlass GitHub repository.
Example
const hgv = hglib.viewer(
document.getElementById('development-demo'),
testViewConfig,
{ bounded: true },
);
Creating a HiGlass component in your React app¶
<HiGlassComponent
options={options}
viewConfig={viewConfig}
/>
Use the HiGlassComponent to create a HiGlass instance in react. The
options prop is the same as explained above.
Example
import { HiGlassComponent } from 'higlass';
const HiGlass = props => <HiGlassComponent
ref={props.onRef}
options={props.options}
viewConfig={props.viewConfig}
/>
export default HiGlass;
Obtaining ordered chromosome info¶
HiGlass provides an API for obtaining information about chromosomes and the order they are listed in a chromSizes file:
import { ChromosomeInfo } from 'higlass';
const chromInfo = ChromosomeInfo(
'http://higlass.io/api/v1/chrom-sizes/?id=Ajn_ttUUQbqgtOD4nOt-IA',
(chromInfo) => { console.log('chromInfo:', chromInfo); });
This will return a data structure with information about the chromosomes listed:
{
chrPositions: {
chr1 : {id: 0, chr: "chr1", pos: 0},
chr2 : {id: 1, chr: "chr2", pos: 249250621} ,
...
},
chromLengths: {
chr1: "249250621",
chr2: "243199373",
...
},
cumPositions: [
{id: 0, chr: "chr1", pos: 0},
{id: 1, chr: "chr2", pos: 249250621},
...
]
}
Convert absolute to chromosomal coordinates:
absPos = 257893;
chromPos = chromInfo.absToChr(absPos);
API Functions¶
-
viewer(element, viewConfig, options)¶ Create a HiGlass component.
In addition to the parameters below, a number of extra options can be passed using the options parameter:
- authToken (string) - An auth token to be included with every tile request
(e.g.
JWT xyz) - bounded (bool) - A boolean specifying whether the component should be sized to fit within the enclosing div [default=false]. If it is false, then the component will grow as needed to fit the tracks within it.
- editable (bool) - Can the layout be changed? If false, the view headers will
be hidden. This can also be specified in the viewconfig using the
editableoption. The value passed here overrides the value in the viewconf. [default=true] - defaultTrackOptions (dict) - Specify a set of default options that will be used for
newly added tracks. These can be broken down into two types: all - affecting all all track types and trackSpecific which will affect only some track types. See the example below for a concrete demonstration.Arguments: - element (Object) – DOM element the HiGlass component should be attached to.
- viewConfig (Object|String) – The viewconfig to load. If this parameter is a string it will be interpreted as a url from which to retrieve the viewconf. If it is a dictionary it will be loaded as is.
- options (Object) – Dictionary of public options. See the description above for a list of available values.
Returns: Object – Newly created HiGlass component.
Examples:
const hgv = hglib.viewer( document.getElementById('development-demo'), testViewConfig, { bounded: true, defaultTrackOptions: { all: { showTooltip: true, }, trackSpecific: { 'heatmap': { showTooltip: false, } } } });
- authToken (string) - An auth token to be included with every tile request
(e.g.
-
reload(target)¶ Reload all or specific tiles for viewId/trackId
Arguments: - target (array) – Should be an array of type ({ viewId: string, trackId: string } | string). If the array is just strings, it’s interpreted as a list of views whose tracks to reload.
-
setViewConfig(newViewConfig, resolveImmediately)¶ Set a new view config to define the layout and data of this component
Arguments: - newViewConfig (obj) – A JSON object that defines the state of the HiGlassComponent
- resolveImmediately (boolean) – If true, the returned promise resolves immediately even if not all data has loaded. This should be set to true, if the new viewconf does not request new data. Default: false.
Returns: Promise – dataLoaded A promise that resolves when all of the data for this viewconfig is loaded. If resolveImmediately is set to true, the promise resolves without waiting for the data to be loaded.
Examples:
const p = hgv.setViewConfig(newViewConfig); p.then(() => { // the initial set of tiles has been loaded });
-
getLocation(viewId="null")¶ Get the current location for a view.
Arguments: - viewId (string) – The id of the view to get the location for
Returns: obj – A an object containing four arrays representing the domains and ranges of the x and y scales of the view.
Examples:
const {xDomain, yDomain, xRange, yRange} = hgv.getLocation('viewId');
-
getMinMaxValue(viewId, trackId, ignoreOffScreenValues=false, ignoreFixedScale=false)¶ Get the minimum and maximum visible values for a given track.
Arguments: - viewId (string) – The id of the view containing the track.
- trackId (string) – The id of the track to query.
- ignoreOffScreenValues (bool) – If
trueonly truly visible values are considered. Otherwise the values of visible tiles are used. Not that considering only the truly visible values results in a roughly 10x slowdown (from 0.1 to 1 millisecond). - ignoreFixedScale (bool) – If
truepotentially fixed scaled values are ignored. I.e., if the absolute range is[1, 18]but you have fixed the output range to[4, 5]you would normally retrieve[4, 5]. Having this option set totrueretrieves the absolute[1, 18]range.
Returns: Array – The minimum and maximum value
Examples:
const [minVal, maxVal] = hgv.getMinMaxValue('myView', 'myTrack');
-
getRangeSelection()¶ Get the current range selection
Returns: Array – The current range selection
-
getTrackObject(viewId, trackId)¶ Return the track’s javascript object. This is useful for subscribing to data events (dataChanged)
Arguments: - viewId (string) – The id of the view containing the track
- trackId (string) – The id of the track within the view
Returns: obj – The HiGlass track object for this track
-
getViewConfig()¶ Retrieve the visible viewconf.
Returns: (Object) A JSON object describing the visible views
Generate a sharable link to the current view config. The url parameter should contain the API endpoint used to export the view link (e.g. ‘http://localhost:8989/api/v1/viewconfs’). If it is not provided, the value is taken from the exportViewUrl value of the viewconf.
Arguments: - url (string) – Custom URL that should point to a higlass server’s view config endpoint, i.e., http://my-higlass-server.com/api/v1/viewconfs/.
Returns: Object – Promise resolving to the link ID and URL.
Examples:
hgv.shareViewConfigAsLink('http://localhost:8989/api/v1/viewconfs') .then((sharedViewConfig) => { const { id, url } = sharedViewConfig; console.log(`Shared view config (ID: ${id}) is available at ${url}`) }) .catch((err) => { console.error('Something did not work. Sorry', err); })
-
zoomToDataExtent(viewUid)¶ Zoom so that the entirety of all the datasets in a view are visible. The passed in
viewUidshould refer to a view which is present. If it doesn’t, an exception will be thrown. Note that if this function is invoked directly after a HiGlass component is created, the information about the visible tilesets will not have been retrieved from the server andzoomToDataExtentwill not work as expected. To ensure that the visible data has been loaded from the server, use thesetViewConfigfunction and placezoomToDataExtentin the promise resolution.Arguments: - viewUid (string) – The view uid of the view to zoom
Examples:
const p = hgv.setViewConfig(newViewConfig); p.then(() => { hgv.zoomToDataExtent('viewUid'); });
-
setViewConfig(newViewConfig, resolveImmediately)¶ Set a new view config to define the layout and data of this component
Arguments: - newViewConfig (obj) – A JSON object that defines the state of the HiGlassComponent
- resolveImmediately (boolean) – If true, the returned promise resolves immediately even if not all data has loaded. This should be set to true, if the new viewconf does not request new data. Default: false.
Returns: Promise – dataLoaded A promise that resolves when all of the data for this viewconfig is loaded. If resolveImmediately is set to true, the promise resolves without waiting for the data to be loaded.
Examples:
const p = hgv.setViewConfig(newViewConfig); p.then(() => { // the initial set of tiles has been loaded });
-
public.zoomTo(viewUid, start1Abs, end1Abs, start2Abs, end2Abs, animateTime)¶ Change the current view port to a certain data location. When
animateTimeis greater than 0, animate the transition. If working with genomic data, a chromosome info file will need to be used in order to calculate “data” coordinates from chromosome coordinates. “Data” coordinates are simply the coordinates as if the chromosomes were placed next to each other.Arguments: - viewUid (string) – The identifier of the view to zoom
- start1Abs (Number) – The x start position
- end1Abs (Number) – The x end position
- start2Abs (Number) – (optional) The y start position. If not specified start1Abs will be used.
- end2Abs (Number) – (optional) The y end position. If not specified end1Abs will be used
- animateTime (Number) – The time to spend zooming to the specified location
Examples:
// Absolute coordinates hgApi.zoomTo('view1', 1000000, 1100000, 2000000, 2100000, 500); // Chromosomal coordinates hglib // Pass in the URL of your chrom sizes .ChromosomeInfo('//s3.amazonaws.com/pkerp/data/hg19/chromSizes.tsv') // Now we can use the chromInfo object to convert .then((chromInfo) => { // Go to PTEN hgApi.zoomTo( viewConfig.views[0].uid, chromInfo.chrToAbs(['chr10', 89596071]), chromInfo.chrToAbs(['chr10', 89758810]), chromInfo.chrToAbs(['chr10', 89596071]), chromInfo.chrToAbs(['chr10', 89758810]), 2500 // Animation time ); }); // Just in case, let us catch errors .catch(error => console.error('Oh boy...', error)) // Using getLocation() for coordinates let firstViewLoc = hgApi.getLocation(oldViewUid); hgApi.zoomTo( viewUid, firstViewLoc["xDomain"][0], firstViewLoc["xDomain"][1], firstViewLoc["yDomain"][0], firstViewLoc["yDomain"][1] );
-
public.zoomToGene(viewUid, geneName, padding, animateTime)¶ Change the current view port to a location near the gene of interest. When
animateTimeis greater than 0, animate the transition.Arguments: - viewUid (string) – The identifier of the view to zoom
- geneName (string) – The name of gene symbol to search
- padding (string) – The padding (base pairs) around a given gene for the navigation
- animateTime (Number) – The time to spend zooming to the specified location
Examples:
// Zoom to the location near 'MYC' hgApi.zoomToGene('view1', 'MYC', 100, 2000);
-
public.suggestGene(viewUid, keyword, callback)¶ Get the list of genes of top match for a given keyword.
Arguments: - viewUid (string) – The id of the view containing the track.
- keyword (string) – The substring of gene name to search.
- callback (function) – A function to be called upon gene list search.
Examples:
hgv.suggestGene('view1', 'MY', (suggestions) => { if(suggestions && suggestions.length > 0) { console.log('Gene suggested', suggestions[0].geneName); console.log('Chromosome', suggestions[0].chr); console.log('Start position', suggestions[0].txStart); console.log('End position', suggestions[0].txEnd); } });
-
exportAsSvg()¶ Get the current view as an SVG. Relies on all the tracks implementing their respective exportAsSVG methods.
Returns: string – An SVG string of the current view.
-
exportAsPngBlobPromise()¶ Get a Promise which returns a Blob containing a PNG for the current view. It’s possible to get string of the PNG bytes from that:
Returns: promise – Examples:
hgApi.exportAsPngBlobPromise().then(function(blob) { var reader = new FileReader(); reader.addEventListener("loadend", function() { var array = new Uint8Array(reader.result.slice(0,8)); console.log(array); console.log(new TextDecoder("iso-8859-2").decode(array)); }); reader.readAsArrayBuffer(blob); });
-
exportAsViewConfString()¶ Export the current view as a Viewconf.
Returns: string – A stringified version of the current viewconf
Generate a sharable link to the current view config. The url parameter should contain the API endpoint used to export the view link (e.g. ‘http://localhost:8989/api/v1/viewconfs’). If it is not provided, the value is taken from the exportViewUrl value of the viewconf.
Arguments: - url (string) – Custom URL that should point to a higlass server’s view config endpoint, i.e., http://my-higlass-server.com/api/v1/viewconfs/.
Returns: Object – Promise resolving to the link ID and URL.
Examples:
hgv.shareViewConfigAsLink('http://localhost:8989/api/v1/viewconfs') .then((sharedViewConfig) => { const { id, url } = sharedViewConfig; console.log(`Shared view config (ID: ${id}) is available at ${url}`) }) .catch((err) => { console.error('Something did not work. Sorry', err); })
-
public.on(event, callback, viewId)¶ Subscribe to events
HiGlass exposes the following event, which one can subscribe to via this method:
- location
- rangeSelection
- viewConfig
- mouseMoveZoom
Event types
click: Returns clicked objects. (Currently only clicks on 1D annotations are captured.){ type: 'annotation', event: { ... }, payload: [230000000, 561000000] }
cursorLocation:Returns an object describing the location under the cursor{ absX: 100, absY: 200, relX: 50, relY: 150, relTrackX: 50, relTrackY: 100, dataX: 10000, dataY: 123456, isFrom2dTrack: false, isFromVerticalTrack: false, }
location:Returns an object describing the visible region{ xDomain: [1347750580.3773856, 1948723324.787681], xRange: [0, 346], yDomain: [1856870481.5391564, 2407472678.0075483], yRange: [0, 317] }
rangeSelection:Returns a BED- (1D) or BEDPE (1d) array of the selected data and genomic range (if chrom-sizes are available)// Global output { dataRange: [...] genomicRange: [...] } // 1D data range [[1218210862, 1528541001], null] // 2D data range [[1218210862, 1528541001], [1218210862, 1528541001]] // 1D or BED-like array [["chr1", 249200621, "chrM", 50000], null] // 2D or BEDPE-like array [["chr1", 249200621, "chr2", 50000], ["chr3", 197972430, "chr4", 50000]]
viewConfig:Returns the current view config (as a string).- This event is published upon interactions including:
- Saving in the view config editor modal.
- Panning and zooming in views, which update view object
initialXDomainandinitialYDomainvalues. - Brushing inviewport-projection-tracks containing nullfromViewUidfields, which update track objectprojectionXDomainandprojectionYDomainvalues.
mouseMoveZoom:Returns the location and data at the mouse cursor’s screen location.{ // Float value of the hovering track data, // Absolute x screen position of the cursor in px absX, // Absolute y screen position of the cursor in px absY, // X screen position of the cursor in px relative to the track extent. relX, // Y screen position of the cursor in px relative to the track extent. relY, // Data x position of the cursor relative to the track's data. dataX, // Data y position of the cursor relative to the track's data. dataY, // Track orientation, i.e., '1d-horizontal', '1d-vertical', or '2d' orientation: '1d-horizontal', // The following properties are only returned when hovering 2D tracks: // Raw Float32Array dataLens, // Dimension of the lens, e.g., 3 (the lens is squared so `3` corresponds // to a 3x3 matrix represented by an array of length 9) dim, // Function for converting the raw data values to rgb values toRgb, // Center position of the data or genomic position (as a BED array) center, // Range of the x data or genomic position (as a BEDPE array) xRange, // Range of the y data or genomic position (as a BEDPE array) yRange, // If `true` `center`, `xRange`, and `yRange` are given in genomic positions isGenomicCoords }
createSVG:Set a callback to obtain the current exported SVG DOM node,- and potentially return a manipulated SVG DOM node.
Arguments: - event (string) – One of the events described below
- callback (function) – A callback to be called when the event occurs
- viewId (string) – The view ID to listen to events
Examples:
let locationListenerId; hgv.on( 'location', location => console.log('Here we are:', location), 'viewId1', listenerId => locationListenerId = listenerId ); const rangeListenerId = hgv.on( 'rangeSelection', range => console.log('Selected', range) ); const viewConfigListenerId = hgv.on( 'viewConfig', range => console.log('Selected', range) ); const mmz = event => console.log('Moved', event); hgv.on('mouseMoveZoom', mmz); const wheelListener = event => console.log('Wheel', event); hgv.on('wheel', wheelListener); hgv.on('createSVG', (svg) => { const circle = document.createElement('circle'); circle.setAttribute('cx', 100); circle.setAttribute('cy', 100); circle.setAttribute('r', 50); circle.setAttribute('fill', 'green'); svg.appendChild(circle); return svg; }); const geneSearchListener = event => { console.log('Gene searched', event.geneSymbol); console.log('Range of the gene', event.range); console.log('Center of the gene', event.centerX); } hgv.on('geneSearch', geneSearchListener);
-
public.off(event, listener, viewId)¶ Cancel a subscription.
Arguments: - event (string) – One of the available events
- listener (function) – The function to unsubscribe
- viewId (string) – The viewId to unsubscribe it from (not strictly necessary) The variables used in the following examples are coming from the examples of
on().
Examples:
hgv.off('location', listener, 'viewId1'); hgv.off('rangeSelection', rangeListener); hgv.off('viewConfig', viewConfigListener); hgv.off('mouseMoveZoom', mmz); hgv.off('wheel', wheelListener); hgv.off('createSVG'); hgv.off('geneSearch', geneSearchListener);
-
setBroadcastMousePositionGlobally(isBroadcastMousePositionGlobally)¶ Enable or disable broadcasting the mouse position globally
Arguments: - isBroadcastMousePositionGlobally (boolean) – If true the mouse position will be broadcasted globally.
-
setShowGlobalMousePosition(isShowGlobalMousePosition)¶ Enable or disable showing the global mouse position
Arguments: - isShowGlobalMousePosition (boolean) – If true the global mouse position will be shown for any track that has options.showMousePosition = true.
-
setGlobalMousePosition(isGlobalMousePosition)¶ - This function is equivalent to calling
- setBroadcastMousePositionGlobally() and setShowGlobalMousePosition().
Arguments: - isGlobalMousePosition (boolean) – If true the global mouse position will be shown and broadcasted.
-
option(key, value)¶ Set or get an option.
Arguments: - key (string) – The name of the option you want get or set
- value (*) – If not undefined, key will be set to value
Returns: obj – When value is undefined the current value of key will be returned.